Institute for Response-Genetics (e.V.)Chairman: Prof. Dr. Hans H. StassenPsychiatric Hospital (KPPP), University of Zurich |

|


|
Master.GEN Version 7.3In order to meet the requirements of complex molecular-genetic studies, we have developed this program package Master.GEN which is based on the experiences from our previous systems Master.EEG for investigations into the structural properties of brain wave patterns, and Master.VOX for investigations into the nonverbal aspects of human speech. For easy use, Master.GEN has been built around a databank, thus facilitating the storage and retrieval of genotype and phenotype data at the different stages of analysis. As a key feature, this program package supports the structural decomposition of genetic diversity by means of adaptive algorithms. Standard statistical analyses, on the other hand, are accessed by interfacing to generally available programs, such as the statistical packages SAS and SPSS, amongst others. Support for Adaptive StrategiesOnce oligogenic configurations of genomic loci have been detected, Neural Network Analysis (NNA) provides powerful tools for modeling pre-specified responses to complex, multidimensional input stimuli. Thus, a classification of patients treated with antidepressants or antipsychotics into early/late/non-responders, for example, can be accomplished using oligogenic configurations of candidate genes as input data. It is the specific advantage of NNA that no causal relationship between stimuli and responses is required, so that experienced users can fit virtually any set of nondegenerate stimuli to any set of responses, provided a sufficiently large and representative set of learning probes is available.
Data Retrieval
Consistency of Genotype Data
Basic Statistics
Principal Components of Genotype Data
Genetic Similarity/Diversity
Simulation of Genotype Data
Structural Analyses
|
|
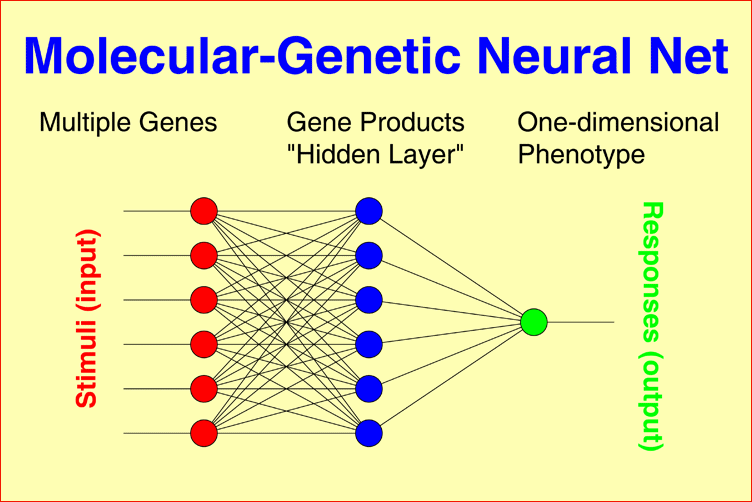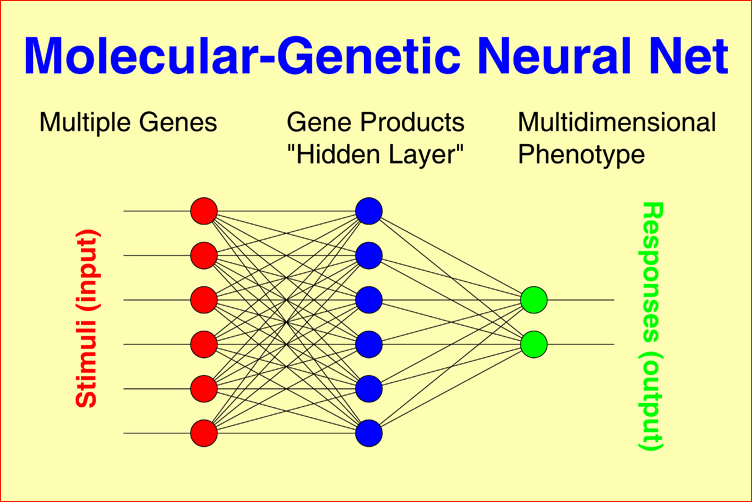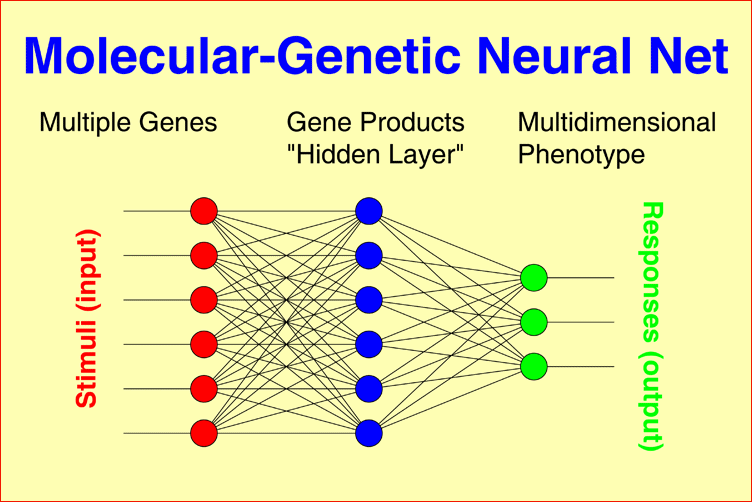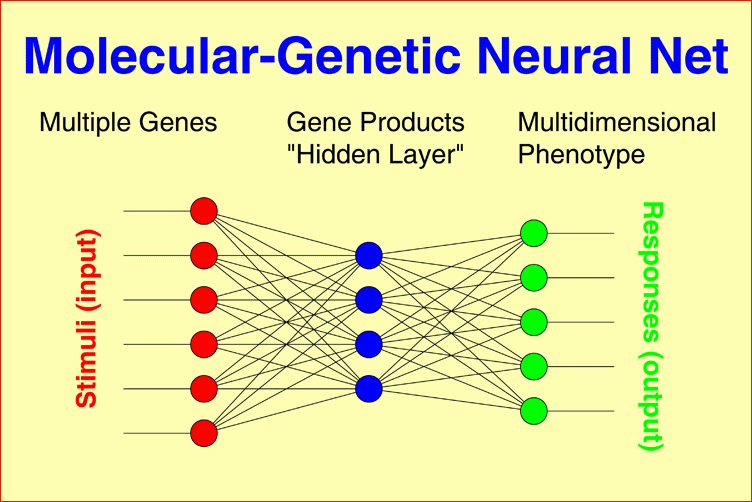
Molecular-genetic Neural Nets may connect multiple genetic factors, as observed in each individual patient, through a layer of gene products to a one-dimensional phenotype, for example, IgM level, Within-pair concordance of monozygotic twins, or time to response to treatment under consideration of interactions between all gene products. The model can easily be generalized to multidimensional phenotypes, for example, the syndrome patterns underlying schizophrenic or bipolar illness. |
|
| [ Mail to Webmaster ] k454910@ifrg.ch |